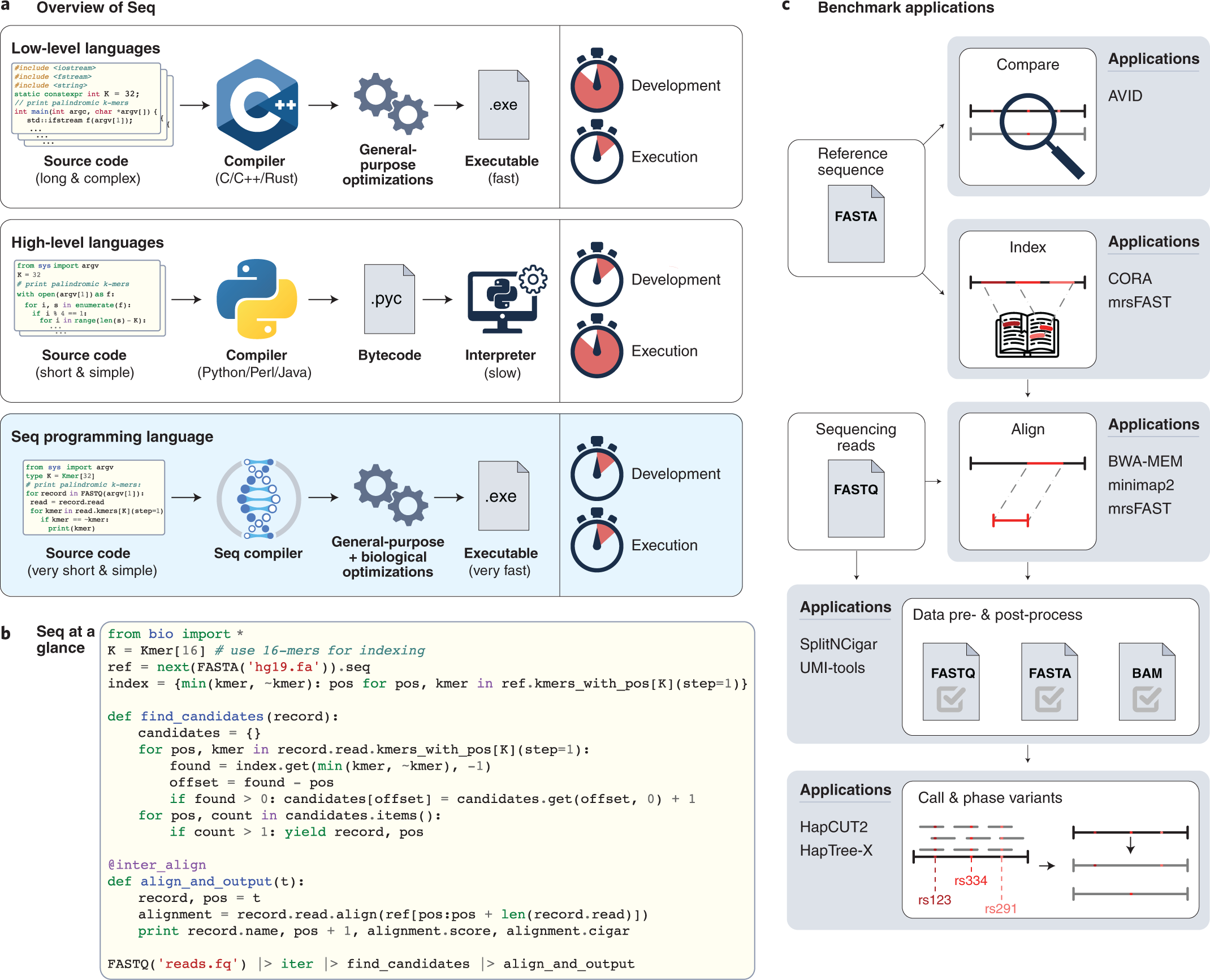
A Python-based programming language for high-performance computational genomics | Nature Biotechnology

A Definitive Guide to Creating Python UDTFs Directly within the Snowflake User Interface - InterWorks
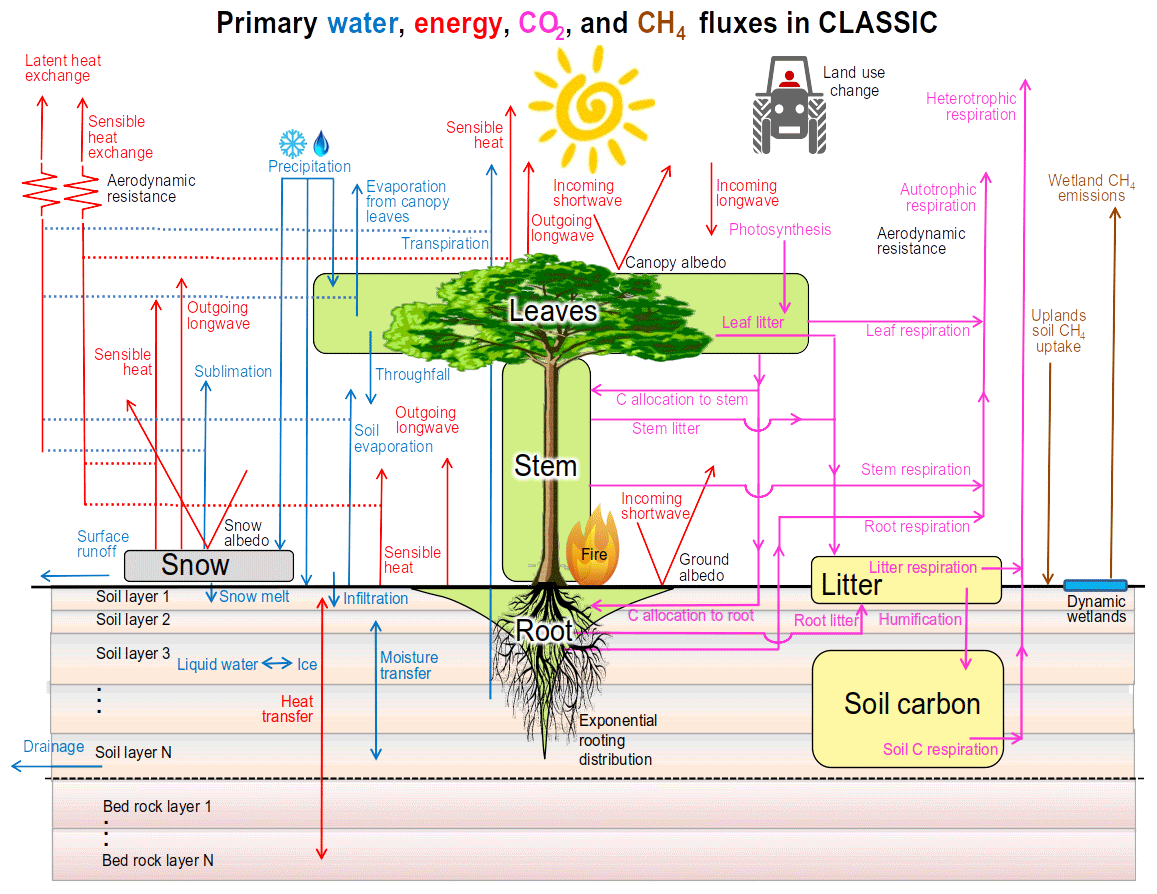
GMD - CLASSIC v1.0: the open-source community successor to the Canadian Land Surface Scheme (CLASS) and the Canadian Terrestrial Ecosystem Model (CTEM) – Part 1: Model framework and site-level performance